B/D HPP
Human Glycoproteomics Initiative
GET INVOLVED: Get in touch at office@hupo.org to learn more
Mission & Goals
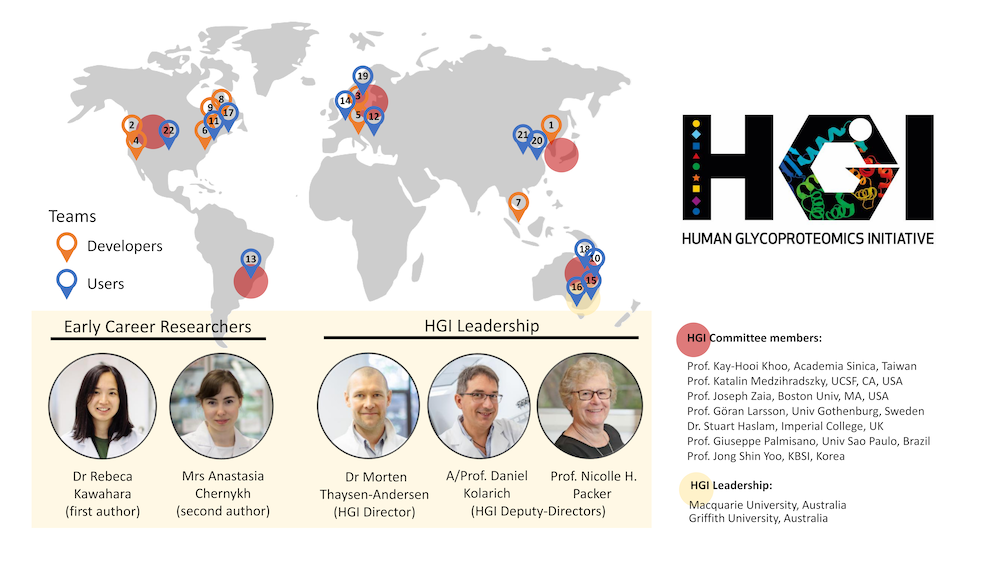
The Human Glycoproteomics Initiative (HGI) was established in 2017 by Distinguished Prof Nicki Packer and Dr Morten Thaysen-Andersen (both Macquarie University, Sydney, Australia). The HGI evolved from the Human Glyco/Proteomics Initiative (HGPI) (2004-2016) headed by Prof N. Taniguchi and Dr H. Narimatsu. A/Prof Daniel Kolarich (Institute for Glycomics, Griffith University, Gold Coast, Australia) joined the HGI leadership team in 2021.
The HGI aims to increase the understanding of the functional significance of the extensive post-translational modification of proteins by glycans. This can only be achieved when the proteomics community has well integrated analytical and informatics tools available to more easily enable the determination of site-, protein-, cell- and tissue-specific glycoform structural heterogeneity in complex biological systems. While other -omics disciplines including genomics and proteomics have matured over past decades, glycoproteomics remains comparatively under-developed limiting our ability to gain insight into the immensely complex, dynamic and functional glycoproteome. Glycoproteomics may therefore be seen as one of the missing pieces of the -omics jigsaw puzzle.
Leadership Information

Chair:
Morten Thaysen-Andersen, PhD
Macquarie University, Sydney, Australia

Co-Chair:
Daniel Kolarich, PhD
Griffith University, Gold Coast, Australia

Co-Chair:
Nicolle H Packer, PhD
Macquarie University, Sydney, Australia
Early Career Researchers:
Rebeca Kawahara, PhD
Macquarie University, Sydney, Australia
Tiago Oliveira, PhD
IMBA, Austrian Academy of Sciences, Vienna, Austria
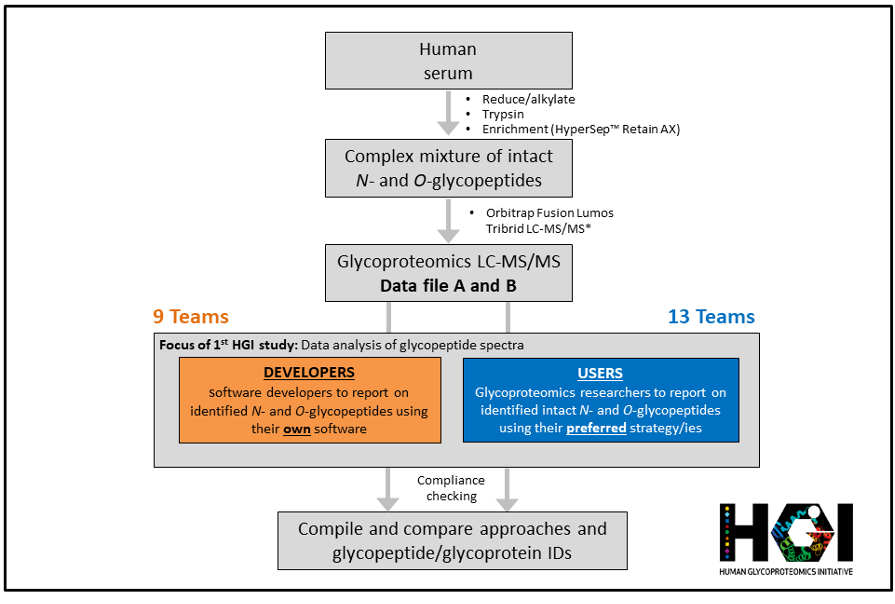
HGI Aims
The HGI is a project/study-centric initiative; our modus operandi is to assemble relevant experts and field-leaders to complete specific studies of particular interest to the community. We seek to progress the field by influencing reporting guidelines (e.g. MIRAGE), nomenclature (SNFG) and experimental recommendations, and by connecting with and bridging into neighbouring disciplines. We aim to promote and draw more attention to our emerging discipline, highlight exciting opportunities and key challenges in the field, and unite like-minded scientists not least to bridge researchers in proteomics and glycomics through dialogue, comparative studies, and open sharing of data, tools and ideas.
The HGI also connects with other Biology/Disease-Human Proteome Project (B/D-HPP) activities in HUPO and permits the exchange of information of mutual interest. The data and information produced by the HGI will be shared widely with the community and will be made available for public, industry, academic research and teaching purposes.