| | | | | |
Chromosome-Centric Human Proteome Project (C-HPP)
|
| |
Mission of the C-HPP
The Human Proteome Project (HPP) of the Human Proteome Organization (HUPO) aims to find high-stringency evidence for all proteins encoded by the human genome, the major splice forms of each protein, mature N- and C-termini, and their major protein post-translational modifications (PTMs). Indeed, one cannot do the best biology and/or medical research without a complete understanding of the parts of the human proteome. Conversely, the international C-HPP teams need the input of biology/disease (B/D) teams to understand the biological context of the parts list. It is a question of focus. A focus on the parts list promotes a measurement- and detail-focused analytical mindset, while a focus on the biological context is focused on outcomes and may ignore individual parts in the context of the overall picture. In conclusion, both mindsets are indivisible parts of the larger HPP initiative.
Questions and Answers on the C-HPP and Missing Proteins: Q & A
Participants (click on flags for more information)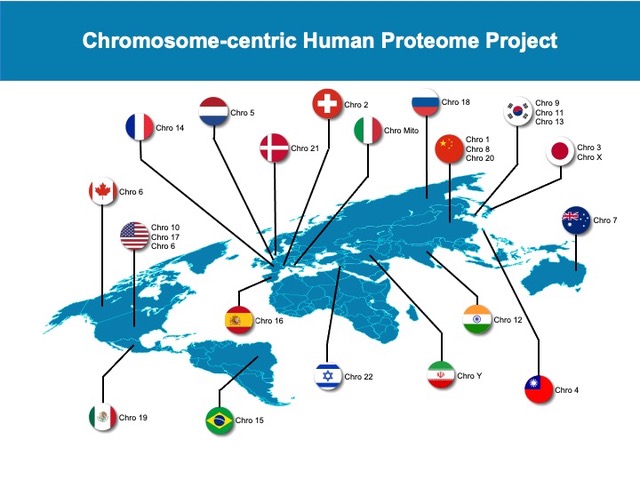
| Chromosome teams | | Chromosome 1: Ping Xu - China | Chromosome 14: Charles Pineau - France
| | Chromosome 2: Vacant - TBD | Chromosome 15: Gilberto Domont - Brazil
| | Chromosome 3: Takeshi Kawamura - Japan | Chromosome 16: Fernando Corrales - Spain
| | Chromosome 4: Yu Ju Chen - Taiwan | Chromosome 17: Gilbert S. Omenn - USA
| | Chromosome 5: Peter Horvatovitch - The Netherlands | Chromosome 18: Elena Ponomarenko - Russia | | Chromosome 6: Rob Moritz - USA/Canada | Chromosome 19: Sergio Encarnacion-Guevera - Mexico | | Chromosome 7: Edouard Nice - Australia | Chromosome 20: Siqi Liu - China
| | Chromosome 8: Gong Zhang - China | Chromosome 21: Frank Schmidt - Qatar | | Chromosome 9: Je-Yoel Cho - Korea | Chromosome 22: Oded Kleifeld - Israel
| | Chromosome 10: Josh Labaer - USA | Chromosome X: Yasushi Ishihama - Japan
| | Chromosome 11: Heeyoun Hwang - Korea | Chromosome Y: Ghasem Hosseini Salekdeh - Iran
| | Chromosome 12: Ravi Siredeshmukh - India | Chromosome Mitochondrial: Andrea Urbani - Italy | | Chromosome 13: Min-Sik Kim - Korea |
|
|
Journal of Proteome Research Virtual issue Celebrates 10 Years of the HPP and C-HPP (October 2020)
2022 C-HPP Executive Committee (EC)
Chair: Christopher M. Overall, Canada (to December 31, 2024)
Co-Chair: Gong Zhang, China (to December 31, 2027) TBA (to December 31, 2028), Co-chair Secretary General: Peter Horvatovich, The Netherlands (to December 31, 2025) Member-at-Large: Sergio Encarnation-Guevara, Mexico (to December 31, 2027)
Member-at-Large: Fernando Corrales, Spain (to December 31, 2025)
Member-at-Large: Gilberto Domont, Brazil (to December 31, 2024) Member-at-Large: Heeyoun Hwang, Republic of Korea (to December 31, 2026)
C-HPP Meetings
The Chromosome Centric Human Proteome Project (C-HPP) announces the 25th C-HPP Workshop on “Accelerating the HPP Grand Challenge” to be held in Algarve, Portugal on Friday, May 13, 2022 following the 2nd Joint Meeting of Spanish, French, and Portuguese Proteomics Societies.
neXt-CP50 Reports
By
analogy to the term “dark proteins” coined to represent structurally
uncharacterized regions, C-HPP investigators have recently adopted the
term “dark proteome” to collectively refer to those proteins for which
we have insufficient
information
on either protein expression, structure, function, or all of these:
They include, for example, MPs (PE2−4), PE5, uPE1 proteins, and any
potential proteins translated from smORF or lncRNAs. When focused on
uPE1 proteins, there are nearly 2,000 proteins which have no functional
information (Paik et al., 2018, JPR, DOI:10.1021/acs.jproteome.8b00383).
On
March 1, 2018, HUPO C-HPP announced launching the neXt-CP50 where CP
stands for “characterization of proteins”. This pilot project aims to
characterize function of 50 uPE1 proteins during ~3 years (2018-2021).
This challenge is to test the feasibility of the functional
characterization of large numbers of dark proteins, 2000 at present the
15 teams are focusing on specific tractable targets that can be
investigated
Of
the C-HPP consortium international teams, 15 from 12 countries joined
this project: Chr 2 (Switzerland), Chr 3 (Japan), Chr 4 (Taiwan), Chr 9,
11, 13 (Korea), Chr 10, Chr 14 (France), Chr 15 (Brazil), Chr 16
(Spain), Chr 17 (USA), Chr 18 (Russia), Chr 19 (Mexico), Chr 20 (China),
and Chr Y (Iran).
C-HPP Papers
- Overall,
C.M. 2020. The HUPO High-stringency Inventory of Humanities Shared
Human Proteome Revealed. Journal of Proteome Research 19, 4,211 – 4,214,
doi/10.1021/acs.jproteome.0c00794.
- Overall, C.M. 2020. The Human Proteome: 90% in the Light — 10% on the Dark Side. Journal of Proteome Research 19, 4,731 – 4,735. https://dx.doi.org/10.1021/acs.jproteome.0c00914
- Omenn,
G.S., Lane, L., Overall, C.M., Cristea, I., Corrales, F., Lindskog, C.,
Paik, Y-K., Van Eyk, J., Liu, S., Snyder, M., Baker, M., Bandeira, N.,
Aebersold, R., Moritz, R., Deutsch, E. 2020. Research on The Human
Proteome Reaches a Major Milestone: >90% of Predicted Human Proteins
Now Credibly Detected, according to the HUPO Human Proteome Project.
Journal of Proteome Research 19, 4,735 – 4,746.
doi/10.1021/acs.jproteome.0c00485.
- Omenn,
Gilbert;Lane, Lydie; Overall,Christopher;Corrales, Fernando;Schwenk,
Jochen;Paik, Young-Ki; Van Eyk, Jennifer;Pennington, Stephen;Snyder,
Michael; Baker, Mark; Deutsch, Eric. Progress on Identifying and
Characterizing the Human Proteome: 2018-2019 Metrics from the HUPO Human
Proteome Project.
- Jang,
K. H., Yoon, H. N., Lee, J., Yi, H. et al., Liver disease-associated
keratin 8 and 18 mutations modulate keratin acetylation and
methylation. FASEB J. 33, 9030-9043 (2019). PMID: 31199680
- Kopylov,
A. T., Ponomarenko, E. A., Ilgisonis, E. V., Pyatnitskiy, M. A. et al.,
200+ Protein Concentrations in Healthy Human Blood Plasma: Targeted
Quantitative SRM SIS Screening of Chromosomes 18, 13, Y, and the
Mitochondrial Chromosome Encoded Proteome. J. Proteome Res. 18, 120-129
(2019). PMID: 30480452
- Paik,
Y. K., Lane, L., Kawamura, T., Chen, Y. J. et al., Launching the C-HPP
neXt-CP50 Pilot Project for Functional Characterization of Identified
Proteins with No Known Function. J. Proteome Res. 17, 4042-4050 (2018).
PMID: 30269496
-
Gilbert
S. Omenn,Lydie Lane,Christopher M. Overall,Fernando J. Corrales, Jochen
M. Schwenk, Young-Ki Paik, Jennifer E. Van Eyk, Siqi Liu, Michael
Snyder, Mark S. Baker, and Eric W. Deutsch. J. Proteome Res. 2018, 17,
4031−4041 Progress on Identifying and Characterizing the Human Proteome:
2018 Metrics from the HUPO Human Proteome Project.
-
Toward
Completion of the Human Proteome Parts List: Progress Uncovering
Proteins That Are Missing or Have Unknown Function and Developing
Analytical Methods. J. Proteome Res. 2018, 17, 4023−4030
-
Theo
Klein, Ulrich Eckhard,Antoine Dufour,†Nestor Solis, and Christopher M.
Overall. Proteolytic CleavageMechanisms, Function, and “Omic”
Approaches for a Near-Ubiquitous Posttranslational Modification. Chem.
Rev. 2018, 118, 1137−1168
- Nikolaus
Fortelny, Christopher M. Overall, Paul Pavlidis & Gabriela V.
Cohen Can we predict protein from mRNA levels? Nature 509, 582–587
(2014); doi:10.1038/nature13319: 27 July 2017 | VOL 547 | NATURE | e 1 9
- Paik,
Y. K., Omenn, G. S., Hancock, W. S., Lane, L. & Overall, C. M.,
Advances in the Chromosome-Centric Human Proteome Project: looking to
the future. Expert Rev Proteomics 14, 1059-1071 (2017). PMID: 29039980
- Paik,
Y. K., Overall, C. M., Deutsch, E. W., Van Eyk, J. E. & Omenn, G.
S., Progress and Future Direction of Chromosome-Centric Human Proteome
Project. J. Proteome Res. 16, 4253-4258 (2017). PMID: 29191025
- Lee,
J. Y., Lee, H. K., Park, G. W., Hwang, H. et al., Characterization of
Site-Specific N-Glycopeptide Isoforms of α-1-Acid Glycoprotein from an
Interlaboratory Study Using LC-MS/MS. J. Proteome Res. 15, 4146-4164
(2016). PMID: 27760464
- Park,
G. W., Hwang, H., Kim, K. H., Lee, J. Y. et al., Integrated Proteomic
Pipeline Using Multiple Search Engines for a Proteogenomic Study with a
Controlled Protein False Discovery Rate. J. Proteome Res. 15, 4082-4090
(2016). PMID: 27537616
-
Eric
W. Deutsch,Christopher M. Overall,Jennifer E. Van Eyk, Mark S.
Baker,Young-Ki Paik,Susan T. Weintraub,Lydie Lane,Lennart Martens,Yves
Vandenbrouck,Ulrike Kusebauch, William S. Hancock, Henning Hermjakob,
Ruedi Aebersold, Robert L. Moritz,and Gilbert S. Omenn. Human Proteome
Project Mass Spectrometry Data Interpretation Guidelines 2.1. (2016) J.
Proteome Res. 2016, 15, 3961−3970
-
Gilbert
S. Omenn, Lydie Lane, Emma K. Lundberg, Ronald C. Beavis, Christopher
M. Overall, and Eric W. Deutsch. Metrics for the Human Proteome Project
2016: Progress on Identifying and Characterizing the Human Proteome,
Including Post-Translational Modifications. J. Proteome Res. 2016, 15,
3951−3960
-
Cho,
J. Y., Lee, H. J., Jeong, S. K., Kim, K. Y. et al., Combination of
Multiple Spectral Libraries Improves the Current Search Methods Used to
Identify Missing Proteins in the Chromosome-Centric Human Proteome
Project. J. Proteome Res. 14, 4959-4966 (2015). PMID: 26330117
-
C.H.
Borchers, J. Kast, L.J. Foster, K.W.M. Siu, C.M. Overall, T.A.
Binkowski,W.H. Hildebrand, A. Scherer, M. Mansoor, P.A. Keowni. The
Human Proteome Organization Chromosome 6 Consortium: Integrating
chromosome-centric and biology/disease driven strategies Journal of
Proteomics 100 (2014) 60 – 67
-
Paik,
Y. K. & Hancock, W. S., Uniting ENCODE with genome-wide
proteomics. Nat. Biotechnol. 30, 1065-1067 (2012). PMID: 23138303
-
Paik,
Y. K., Jeong, S. K., Omenn, G. S., Uhlen, M. et al., The
Chromosome-Centric Human Proteome Project for cataloging proteins
encoded in the genome. Nat. Biotechnol. 30, 221-223 (2012). PMID:
22398612
-
Na,
K., Lee, M. J., Jeong, H. J., Kim, H. & Paik, Y. K., Differential
gel-based proteomic approach for cancer biomarker discovery using human
plasma. Methods Mol. Biol. 854, 223-237 (2012). PMID: 22311764
-
Legrain,
P., Aebersold, R., Archakov, A., Bairoch, A. et al., The human proteome
project: current state and future direction. Mol. Cell Proteomics 10,
M111.009993 (2011). PMID: 21742803
| | |
|