By Chia-Feng Tsai & Yasushi Ishihama, Kyoto University
Dysregulation of cellular signaling based on protein phosphorylation is closely linked to pathogenesis of human diseases and therapeutic strategies to control the phospho-signaling have been accepted to develop molecular-targeting drugs for cancer. MS-based quantitative phosphoproteomic approaches have been widely used to quantify over 10,000 phosphorylation sites by utilizing strong cation exchange chromatography, hydrophilic interaction chromatography or basic-pH reversed-phase chromatography to fractionate the complex samples. However, such extensive fractionation approaches require longer LC-MS measurement time as well as tedious pretreatment steps, resulting in reduced throughput and low reproducibility. Besides, these approaches cannot be applied to primary cells of rare tissues or clinical biopsy samples due to the limited starting materials.
To address these issues, Humphrey et al. (from Matthias Mann’s group) at Max Planck Institute of Biochemistry have developed a streamlined phosphoproteomics workflow called “EasyPhos” which has been designed as a high throughput and simplified workflow to study time-resolved phosphorylation alteration in vivo without any pre-fractionation strategy. They used trifluoroethanol for the digestion buffer which allows bypassing the peptide desalting step before phosphopeptide enrichment. This protocol can reduce the potential sample loss during the desalting process implemented in the conventional protocols. Besides, this simplified procedure can be expanded to a 96-well plate format to increase the throughput of phosphopeptide enrichment. The combination of this workflow with Q Exactive benchtop Orbitrap mass spectrometer allows monitoring of more than 10,000 phosphorylation sites from a mouse cell line by single-shot LC-MS/MS analysis with 2hr gradient. The applicability of this parallelized EasyPhos workflow has been demonstrated on the analysis of liver phosphoproteomes at different time points (early and intermediate) in fasted mice under insulin exposure. Up to 31,605 phosphopeptides (25,507 phosphorylation sites belong to class 1) from 6,138 phosphoproteins were identified from 91 biologically distinct liver tissues by the high throughput 96-well EasyPhos assay. Importantly, at least six biological replicate analyses (separate mice) per sample for each time point provide a highly statistical power to illustrate time-resolved maps of insulin signaling. Moreover, these dynamics datasets illuminate not only the insulin-mediated signaling network but also the signaling cascade from cell surface to the nucleus within 1 min in vivo. This rapid and high throughput EasyPhos workflow will facilitate to accumulate the knowledges of cellular signaling dynamics under physiological or pathological regulation.
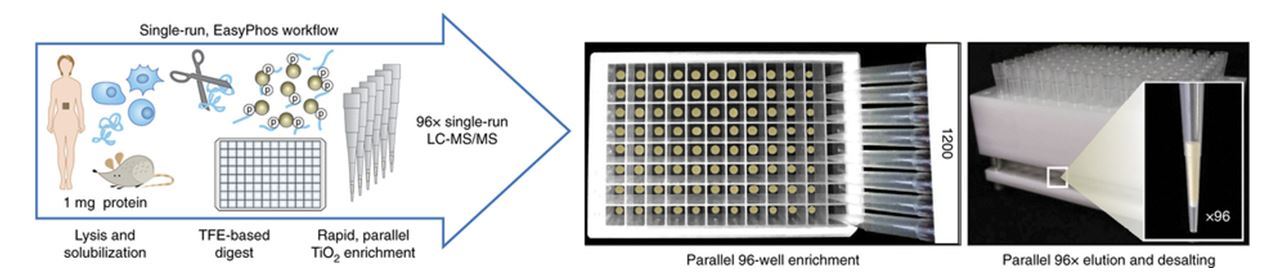 Figure reprinted with permission of the Nature Publishing Group
Figure reprinted with permission of the Nature Publishing Group
This study was reported in the journal of Nature Biotechnology on August 17, 2015.
Link: http://www.nature.com/nbt/journal/v33/n9/abs/nbt.3327.html


.png)